According to reports, it takes ~10 years and $2.6 Billion for a new drug to complete its journey from initial discovery to its market launch. Given the timeline and the resources at stake, you want to make sure you extract maximum ROI from the drug.
The logical step would be to get a patent on your drug so that you can eliminate competition, and monopolize the market. The next question that arises is – How to ensure that this drug is novel and free to launch?
The long and short of it is – You will need a detailed analysis of patent and non-patent literature to check what structure variations (Markush claims/structures) are already present in the prior art, and then decide your patent filing strategy accordingly.
However, searching for Markush/chemical structures is often a complicated endeavor.
Chemical/Markush Structure searching poses several challenges. Let me walk you through some of the challenges.
Markush structures in patents are broad and difficult to interpret as patent applicants try to cover as many functionally equivalent variants as possible.
A typical Markush structure is shown in the following claim.

The listed structure includes variables (R1, R2, X, and R3) at specific positions that create implied structural variations. This means that any of the following chemical moieties and combinations thereof at those listed positions are included in the patent’s claims. Hence, it requires analysis of both structure diagrams and text, and of semantic relationships between them.
The sheer complexity and volume involved make finding specific structures time-consuming. It requires experience and sophisticated search tools to reliably access the necessary information.
Besides the bulkiness of Markush structures, additional hindrances are there in performing efficient chemical patent search like – the presence of stereoisomers, isomers, and the charged moiety of a particular chemical structure.
Also, some patents which claim only the Markush structure of chemical compounds and are silent about their common or trade names can be missed by doing only keyword-based searching.
There are other challenges too but we have the culture of overcoming obstacles and finding prior art. We don’t say this lightly, “if it exists, we will find it.” We have previously illustrated in our blog cases where we overcame all the odds to find prior art and achieve client satisfaction. We approach search projects not as vendors, but as search partners, which means we leave no stone unturned in finding that critical piece of prior art for you, irrespective of the kind of search.
Let me take the example of a recent case to better illustrate this.
Case Study: How do we tackle the challenges of Markush structures searching?
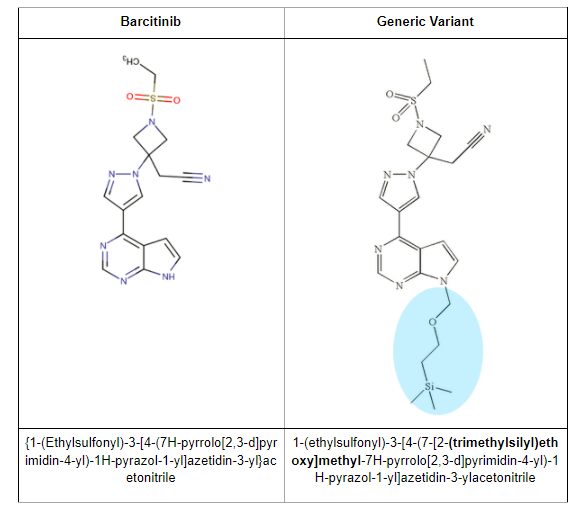
Some time ago, we received a request from Kevin, who is a senior attorney at one of the top US law firms. Kevin’s client was one of the top generic manufacturers in the US. His client wanted to launch a structure variant ((trimethylsilyl)ethoxy]methyl variable) of rheumatoid arthritis drug baricitinib.
They also wanted to file a patent on this. But before going ahead, they wanted to know what kind of prior art already existed for it.
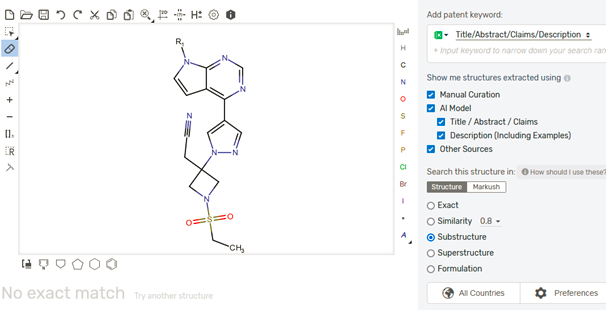
To elaborate a bit, this is the variant they’re trying to launch:
 How do we approach structure searches for such cases?
How do we approach structure searches for such cases?
We initiated our search with an exact structure search on free databases like ChemSpider and Surechembl to capture low-hanging fruits first. This is the best approach if you want to locate results quickly. However, in this case, nothing relevant surfaced.
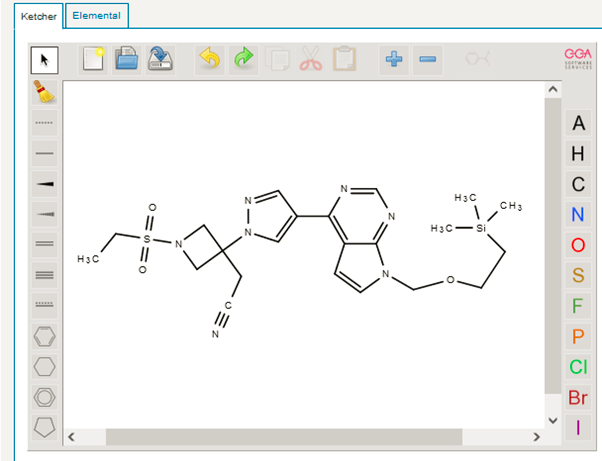
After the analysis on one databases, it was time to dig deeper. Starting with broad search always help. We then next moved our search to other sophisticated paid databases having more coverage like STNext (CAS registry and MARPAT) and Patsnap chemical. There, we executed (sub)structure searching, where we ran the basic motif/scaffold of the structure and tried to locate all the variants and see whether our one was already there or not.
Besides structure searching, we also perform keyword-based searching to ensure comprehensiveness. So, we hunt for IUPAC names or CAS names, common names, trade names, and all possible names of the chemical to find relevant patents from the database. We also gathered INN InchI, InchIkeys, and Smiles for the variant.
In such cases, we also execute searches with different spellings of chemical names. The idea is to not miss any existing prior art just because we missed covering other spelling variations of the chemical names.
For this case, we collected the following details and executed our search accordingly.
| IUPAC names (different spelling and writing style)
| {1-(Ethylsulfonyl)-3-[4-(7-{[2-(trimethylsilyl)ethoxy]methyl}-7H-pyrrolo[2,3-d]pyrimidin-4-yl)-1H-pyrazol-1-yl]-3-azetidinyl}acetonitrile |
| {1-(Ethylsulfonyl)-3-[4-(7-{[2-(trimethylsilyl)ethoxy]methyl}-7H-pyrrolo[2,3-d]pyrimidin-4-yl)-1H-pyrazol-1-yl]-3-azetidinyl}acetonitrile | |
| {1-(Ethylsulfonyl)-3-[4-(7-{[2-(trimethylsilyl)ethoxy]methyl}-7H-pyrrolo[2,3-d]pyrimidin-4-yl)-1H-pyrazol-1-yl]-3-azetidinyl}acetonitrile | |
| {1-(Éthylsulfonyl)-3-[4-(7-{[2-(triméthylsilyl)éthoxy]méthyl}-7H-pyrrolo[2,3-d]pyrimidin-4-yl)-1H-pyrazol-1-yl]-3-azétidinyl}acetonitrile | |
| 2-(1-(ethylsulfonyl)-3-(4-(7-((2-(trimethylsilyl)ethoxy)methyl)-7H-pyrrolo[2,3-d]pyrimidin-4-yl)-1H-pyrazol-1-yl)azetidin-3-yl)acetonitrile | |
| 3-Azetidineacetonitrile, 1-(ethylsulfonyl)-3-[4-[7-[[2-(trimethylsilyl)ethoxy]methyl]-7H-pyrrolo[2,3-d]pyrimidin-4-yl]-1H-pyrazol-1-yl]- | |
| SMILES | N#CCC1(N2N=CC(C=3C4=C(N(COCC[Si](C)(C)C)C=C4)N=CN3)=C2)CN(S(=O)(CC)=O)C1 CCS(=O)(=O)N1CC(C1)(CC#N)N1C=C(C=N1)C1N=CN=C2C=1C=CN2COCC[Si](C)(C)C |
| Std. InChi | InChI=1S/C22H31N7O3SSi/c1-5-33(30,31)28-14-22(15-28,7-8-23)29-13-18(12-26-29)20-19-6-9-27(21(19)25-16-24-20)17-32-10-11-34(2,3)4/h6,9,12-13,16H,5,7,10-11,14-15,17H2,1-4H3 |
| Std. InChIKey | WUWJJJDMZWZKQS-UHFFFAOYSA-N |
The idea was to make sure we conducted a comprehensive search by implementing all the above-mentioned steps. We also manually scrutinized the documents because we understand that the Markush structure search needs expertise in correlating the diagram with the text specified in the patents.
After a very exhaustive comprehensive search, we concluded there was nothing similar to the drug we were looking for. We shared this finding and the approach with Kevin, and he felt quite confident and satisfied with the comprehensiveness of the analysis. With the same confidence, he helped his end client to file a patent for this drug variation too. It was a win-win situation for Kevin and his end client.
We also highlighted a few key patents that could be risky for Kevin’s end client. This information generally helps mitigate the risk before launching the product. A win for the client and we get extra brownie points for customer satisfaction.
Concluding Notes
Chemical structures are described using a variety of different naming conventions, or simply by drawing their chemical/Markush structure. Thus it became notoriously difficult to find relevant chemical/Markush structure patents in the fields of chemistry or pharmaceutical sciences.
We do understand that searching for accurate patented chemical compounds is a daunting task, especially if a proper chemical structure search strategy is not in place.
We thus approach this kind of search with a proper plan and cover all the approaches like structure-based searching (exact and substructure) and even perform keyword-based searching to make sure that we do not miss anything.
Loved what you read and want your help for your next project that involves chemical structure search? Send us a message and we will get back to you to discuss the deets.

Authored by: Divya Goyal, Pharma Team.