Imagine this – You are a leading biopharmaceutical player whose R&D team has developed a novel product. Before launching the product, you want to make sure your product is not infringing on anyone else’s patents. There are various costs involved in manufacturing and marketing this product, and you do not want to get entwined in infringement litigation. So you make the wise move – You ask a vendor to conduct a Freedom to operate analysis on your product.
Now there’s this thing about Freedom to operate searches — If you miss any single relevant patent in the analysis, it can pose fatal repercussions. You would want to make sure your vendor does a very comprehensive search and misses nothing. This is why you must choose the right search vendor partner. This is the part where I would pitch our services, but we will save that for later. I believe more in Show Vs. Tell.
Recently, we were contacted by Sam, who is the lead IP counsel at one of the leading pharmaceutical companies in the US. Their R&D team developed an antibody and Sam wanted to make sure they were not infringing on anyone’s IP and asked us to conduct a freedom to operate search.
In cases that involve sequences like genes and proteins, the FTO search is done quite differently. The process involves a combination of both sequence searching and keyword searching with sequence searching being the most crucial step in the process.
Sequence searching could lead to answers once you overcome the roadblocks
Before we start, here’s something you should know about sequence searching.
Sequence searching is a whole different ball game that requires specific databases and technical expertise to search and interpret sequence alignments. Because in sequence searches, you are expected to report not only exact matches but fragments of identities so that the mutants with a few changes in sequence can be detected.
These lead to a whole world of complexities like:
- Not just 100% matches are relevant: for example – in the case of Etanercept (a fusion protein made up of 2 sequences), the sequence specified in US5395760A which only matches the one sequence (i.e. matches with the first 235 amino acids of the complete 467 amino acid sequence of Etanercept) is a bang-on hit from FTO perspective.
| Etanercept | Prior-art – US5395760A |
|
The fusion protein is made up of 2 parts where
Length – 467 Amino acid 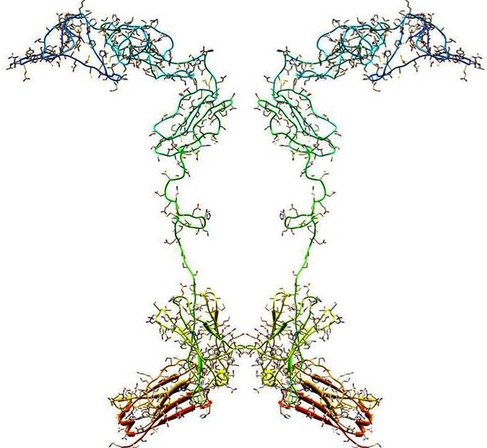
|
Claimed a protein sequence that binds to TNFR. Length – 235 amino acid It matches just half of the etanercept (or I can say, alignment percent identity is only 50%). But still, we can’t miss such a highly relevant patent from the FTO perspective. |
- Not all 100% matches are relevant: For instance, if you are looking for probes, primers, or CDRs, then the query sequence is too short (perhaps 25 residues). Then, there are highly likely chances that these small sequences match by chance in multiple long sequences. But, those long sequences might have different functionality to what we are looking for. In such cases, the alignment percent identity may be useful but the subject % identity is even more useful.
For example — Once we were searching for a CDR, a functional part of an antibody targeting CD28. The CDR has a small length, PGGITTS). We got a 100% match of this CDR with a portion of antibody targeting HER3 (US20190023796A1) as specified below. Although it is a 100% query identity, these kinds of by chance matches are not relevant.
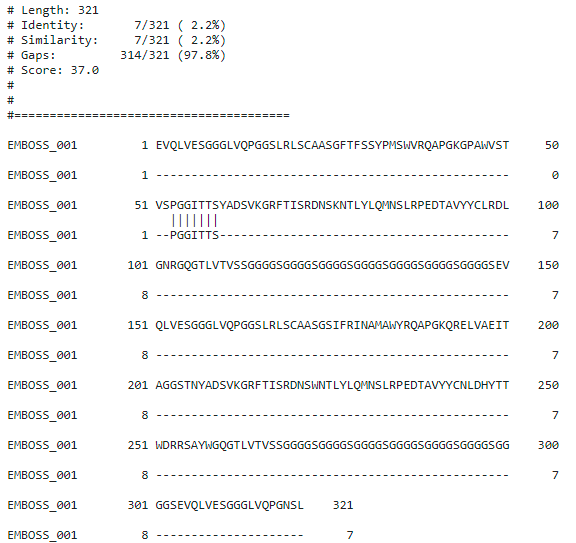
A good searcher would keep these complexities in mind and plan the search accordingly. Back to our show vs. tell. Let me show you how we approach these kinds of searches.
How we conducted FTO for an antibody sequence that binds CD19 to cure cancer?
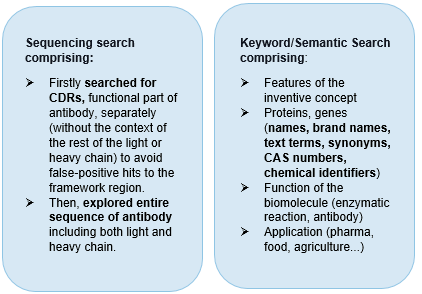
The below visual is a quick depiction of how we plan the whole search. This is a step-by-step process that has sequence search at its core and keyword search for comprehensiveness.
We start with CDR Sequence searching – It is a fact that all antibodies have significant sequence homology to each other since they all have the Immunoglobulin (Ig) domain. So, the chances of getting irrelevant noise are quite high.
Thus, in this case, we searched for CDRs (functional part of antibody) separately (without the context of the rest of the light or heavy chain) to avoid false-positive hits to the framework region.
We started the analysis with 100% seq identity for these CDRs because it is a well-known fact that even a single amino acid change in the CDR might affect the binding to the target. And, because of the small length of CDRs, there surfaced a significant number of patents claiming individual CDRs. But, no single patent disclosed a combination of 3 CDRs.
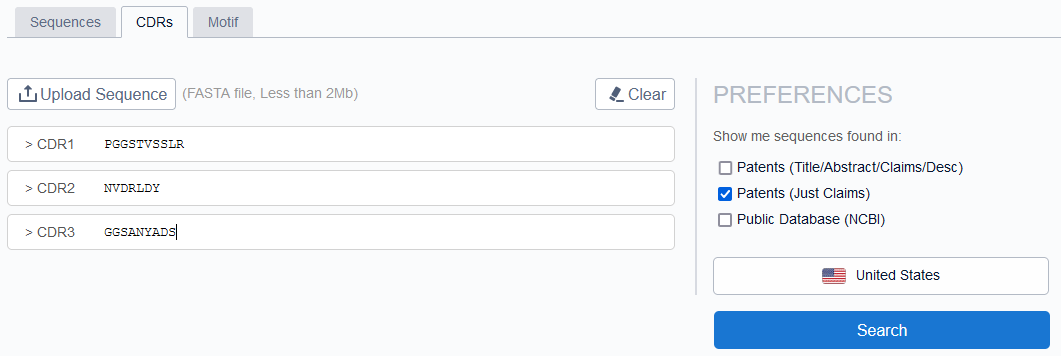
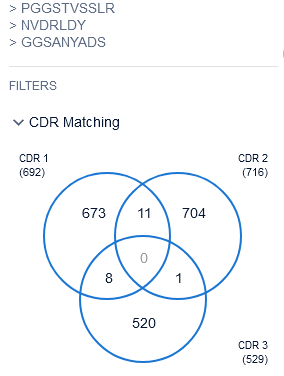
We didn’t stop here. We then executed sequence search by an entire heavy and light chain as well with 85% sequence similarity cut-off. And reviewed them in detail to see whether those sequences belong to the same functionality property (targeting the same antigen) or not.
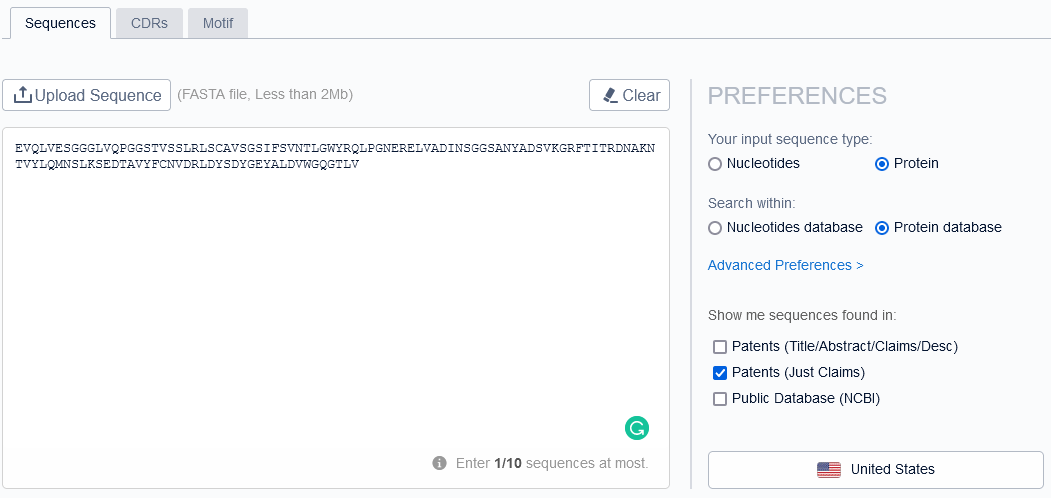
From sequence searching, nothing relevant came up, so we moved our gears to the keyword/semantic-based search. The reason being, in FTO — references that claim functional properties of antibodies (an antibody targeting CD19 antigen) broadly without any sequence constraint would be relevant.
Semantic search is broader than a keyword search but false hits are much higher. But to leave no stone unturned, we always perform a semantic search as well.
From keyword-based searching, we got some US patents claiming broadly antibodies targeting CD19 without the sequence restriction.
We shared these insights with Sam and suggested he gets a license to these patents before launching the product in the market. Sam was pretty satisfied with the quality of the search and shared this feedback:
“When it comes to sequence searches, I have my doubts whether a vendor could perform a comprehensive search considering all the nuances. But you guys did a good job at it. Kudos! “
Concluding Notes
A major roadblock in conducting sequence-based FTO searches includes scattered information across multiple databases, expertise in reducing database noise using sequence-based parameters, lack of sequence analysis expertise combined with claim interpretation skills. It is no wonder that biologics patent searching can often be reminiscent of looking for an ever-smaller needle in an even bigger haystack.
However, successes in sequence searching can be attained by combining various methods like keyword, sequence search, and semantic or concept search. These searches executed by skilled searchers who have an investigative approach towards conducting searches would definitely get you the right results, and void you of unwanted litigation.
Relevant cases where above mentioned strategies can be applicable:
- Nippon Shinyaku.,Ltd. v. Sarepta Therapeutics, Inc. PTAB 1:21-cv-01015
- TwinStrand Biosciences, Inc. et al v. Guardant Health, Inc. 1:21-cv-01126
- Illumina, Inc. v. APExBIO Technology LLC 4:21-cv-02611
Have a sequence search that you need help with? Let’s talk about it. Use the form below and tell us about your requirement and our team will reach out to you in no time:
Authored by: Divya Goyal, Pharma.